
Nicolás Molla
Musician, composer, and music producer. He has created music for film, advertising, and social projects, and now works as an independent producer in his own studio.
nicomolla.com →A fusion of DNA structure, its interaction with the biological environment, and musical composition. An invitation to hear the molecule of life through physical properties extracted from simulations and turned into audible signals.
Sonification is a way of turning data into sound so we can understand it better. Instead of displaying information in graphs or tables, it is transformed into acoustic signals that we can hear. In this way, what would normally be a series of numbers or measurements becomes a “soundscape” that reflects how a phenomenon, experiment, or model behaves.
This process is not automatic; someone has to decide which data will be transformed and how they will sound. For example, a higher sensor reading can be turned into a higher pitch, or a sudden change in a measurement can be heard as a strike or a shift in rhythm. In this way, sonification opens up a new path for exploring, interpreting, and communicating information by taking advantage of our natural ability to recognize patterns in what we hear.

There are also more poetic examples of sonification, such as the approach developed by NASA to let us hear distant galaxies (nasa.gov/marshall):
Molecular sonification takes biological data — protein sequences, gene expression patterns, DNA dynamics, folding trajectories — and maps them onto sound: pitch, rhythm, timbre, instrumentation. The goal is not just to listen to a molecule but to use our ear as an analytical tool, surfacing patterns that are hard to spot on a chart. Below are some of the most representative attempts published in recent years.
Music from protein sequences, with musicality enhanced through a computer program that learns from Chopin.
Conversion of amino acid sequences in proteins into classical music: a search for auditory patterns.
A musical approach to the interpretation of gene expression data using neuroblastoma cell lines.
“Despite the filtering and rearrangement of the probe sets, the resulting melodies in the examples presented are quite abstract, and their evocative potential is difficult to predict. It seems likely that familiarity with such melodies would be achieved more quickly if dissonances from familiar melodies were heard.” (sic)
Musical patterns for comparative epigenomics.
SNARE Dance: a musical interpretation of Atg9 transport to the tubulovesicular cluster.
“After assigning instruments to each protein score, we went on to combine the individual scores into a final orchestration.” (sic)
Hydrogen-bond heterogeneity correlates with transition-state passage time in protein folding.
Molecular dynamics simulations are computer simulations that make it possible to observe how the molecules that make up life — proteins, DNA, RNA — move and change over time. They work by applying the laws of physics to each atom, allowing us to follow their trajectories as if we had a virtual microscope capable of seeing at the atomic level and in slow motion.
These simulations are extremely useful because they allow us to explore phenomena that are impossible to observe directly in the lab, such as exactly how a DNA sequence bends, folds, or becomes more rigid depending on the combination of letters (bases) that make it up. Thanks to this approach, it has become clear that the physical properties of DNA — flexibility, rigidity, and tendency to bend — depend strongly on its sequence.
A key role in this progress has been played, and continues to be played, by the Ascona B-DNA Consortium (ABC), an international collaboration of researchers that has been generating DNA simulations since the early 2000s, establishing standards and databases that are now essential references in the field. DansLab has been part of the ABC Consortium since 2014 and was the most recent organizer of the ABC conference held in April 2023 in Ascona, Switzerland.
Simulated DNA sequences come from the miniABC library provided by the ABC Consortium. They contain 136 unique tetranucleotide combinations made up of the four DNA letters (A, C, G, T) — every possible 4-letter context that can appear in a DNA strand.
From those simulations we extracted a theoretical framework to compute DNA–K⁺ interactions and the concentration of potassium ions in the major and minor grooves. When K⁺ ions sit inside a groove they leave a measurable kinetic and energetic signature that we then map onto sound.
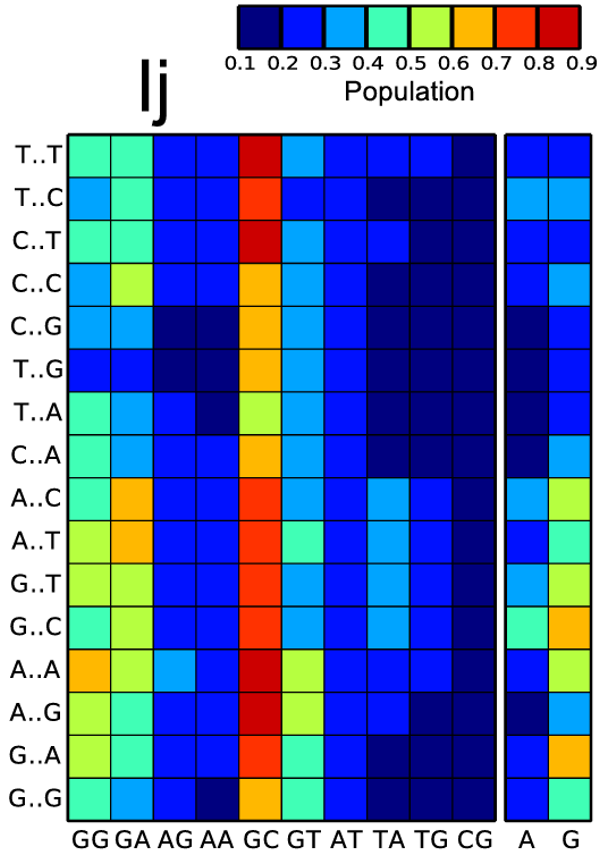


Realizing that sonifications that are difficult or fatiguing to listen to will be less successful, some valiant attempts have been made to incorporate some elements of composition into the sound mappings. As music is designed to engage and hold the listener's interest, surely a sonification that is more musical will be better than one that is not. Unfortunately, sonifications purportedly designed to be musical are often still fatiguing or unengaging. Conversely, the goal of communicating essential information can be masked in the effort to achieve a stronger musical expression.
Trying to follow the balance between data and composition described by Vickers, we transformed the interaction between DNA and potassium cations (K⁺) into music.
For all possible four-letter sequences, the interaction in the major and minor grooves of DNA was measured. The groove-interaction frequencies were multiplied by a factor to bring them into the human audible range. The resulting values were then rounded by mapping the frequencies to the nearest note in the tempered scale.
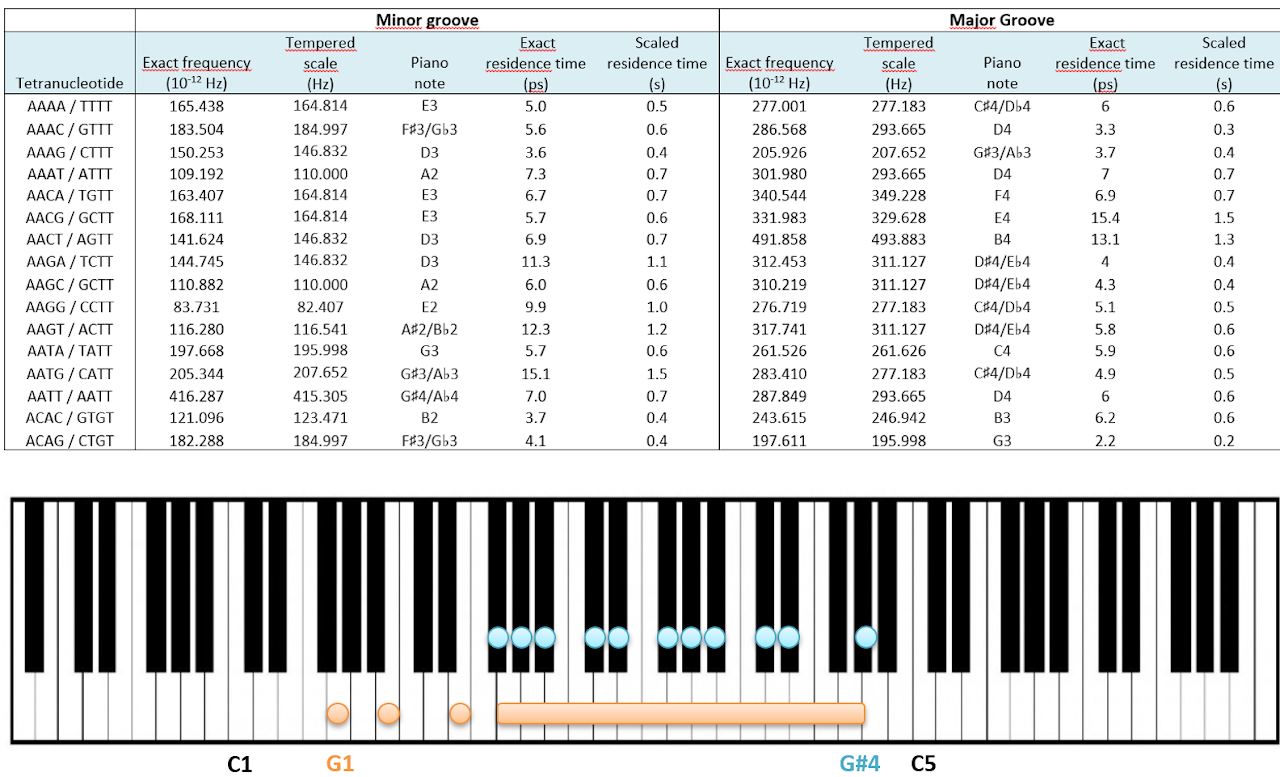
As a pilot test, the 13 miniABC sequences were joined into a single 234-letter sequence (A, C, G and T) and turned into music for piano and violin. Red notes represent DNA–K⁺ interactions in the minor groove, blue notes the major groove, and black notes are part of the musical composition.
Listen on YouTube
An interactive player that lets you type a DNA sequence and hear the music produced by our algorithm. Choose your bases and let the molecule speak.

Musician, composer, and music producer. He has created music for film, advertising, and social projects, and now works as an independent producer in his own studio.
nicomolla.com →
Researcher, teacher, and science communicator. International expert in nucleic acid structure (DNA and RNA) and in computational chemistry, molecular modeling, simulations, and structural bioinformatics.